|
|
|
|
|
|
| Michael J Hendzel, Ph.D. | Principal Investigator |
| Darin McDonald, M.Sc. | Lab Manager |
| Ajit Sharma, Ph.D. | Research Associate |
|
Hilmar Strickfaden, Ph.D.
|
Research Associate |
| Hilmar Strickfaden, Ph.D. | Research Associate |
Approximately 40% of Canadians are expected to have cancer at some point in their lives and approximately 25% of Canadians are expected to die of cancer. It is only recently that modern biology has begun to impact upon the clinical treatment of cancer patients. Despite early successes in so-called rationally designed therapy, much of the fundamental biology that provides the foundation for rationally designed therapy remains to be discovered and characterized. My research laboratory investigates the basic biology of the genome and the cell nucleus, which houses the genome. The maintenance of genome stability (mechanisms that ensure the faithful transmission of chromosome number of sequence content), the regulation of DNA double strand break repair, and the regulation of the genome through epigenetic mechanisms are being studied at the level of single cells with the objective of identifying mechanisms that have the potential to be translated into novel rationale therapies. We are currently funded by the Canadian Institutes of Health Research
|
Current Research Taking Place in Our Laboratory |
|
|
|
Genome Stability and Epigenetics in Cancer. Epigenetics describes the coding for information in the genome that is important in the control of cell phenotype but is not directly encoded in the DNA. It is now appreciated that 50% or more of the cumulative changes in a cell necessary to convert it into a cancer cell is epigenetic, rather than genetic, in origin. We are currently studying how epigenetic modifications of chromatin function in organizing and regulating the genome during interphase and mitosis. In collaboration with Drs. Robert Campbell, Gordon Chan, J.B. Rattner, and Alan Underhill, we are developing reagents and drugs to exploit the requirement for histone methylations in maintaining genome stability. Read more about this research. |
|
|
DNA Damage and DNA Damage Repair. Many current cancer therapies, including radiation therapy, kill cancer cells by introducing double-strand breaks in the DNA. We have found a novel function for nuclear actin in repairing DNA double-strand breaks and are working to identify novel actin-related targets to enhance the effectiveness of radiation therapy. In collaboration with Dr. Joan Turner, we are investigating how changes in genome organization alter the ability of cells to repair radiation-induced double-strand breaks (and thereby fail to kill cells). We are also collaborating with Drs. Guy Poirier and Jean-Yves Masson to determine how cells sense and signal the existence of DNA double-strand breaks. Collectively, these studies have the objective of identifying mechanisms to improve cancer therapies that work through the introduction of double-strand breaks in the DNA. Read more about this research. |
|
|
Nuclear Dynamics and Genome Regulation. One of the frontiers in cell and cancer cell biology is to determine the dynamic properties of important cellular proteins inside of living cells. These studies complement genomics and proteomics studies by providing the details of the spatial and temporal regulation that takes place inside of living cells. Our laboratory is particularly interested in the dynamics of nuclear proteins that take place in the orchestration of genome regulation, chromatin structure, and genome repair. In collaboration with Dr. John Th'ng, we are studying the regulation of the histone H1 protein during cellular transformation and terminal differentiation. We are also characterzing the basic physical properties of the living nucleus in order to understand how these properties influence the regulation of the genome in normal and cancer cells. This is transdisciplinary work involving collaborations with Dr. Gerda de Vries (Mathematics) and Dr. Jack Tuszynski (Biophysics and Computational Biology). Read more. |




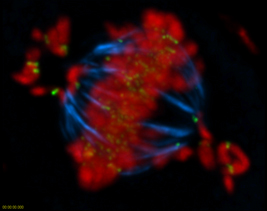 A HeLa cell treated with a protein
methylation inhibitor for 2 hours. The treatment
results in a failure to properly align the chromosomes
during metaphase
A HeLa cell treated with a protein
methylation inhibitor for 2 hours. The treatment
results in a failure to properly align the chromosomes
during metaphase A pair of mouse embryonic fibroblast
cells stained with an antibody recognizing
phosphorylated histone H2AX. The cells have been
exposed to ionizing radiation prior to staining
generating the charactgeristic "γH2AX"
foci at sites of DNA damage.
A pair of mouse embryonic fibroblast
cells stained with an antibody recognizing
phosphorylated histone H2AX. The cells have been
exposed to ionizing radiation prior to staining
generating the charactgeristic "γH2AX"
foci at sites of DNA damage.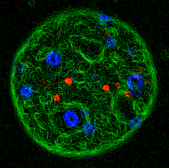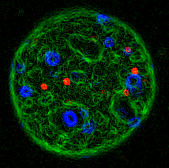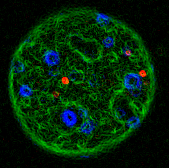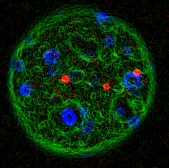 A mouse fibroblast stained for DNA
(blue), transfected with SC35-GFP (a fluorescently
tagged RNA splicing factor) and microinjected with 40 nm
beads. A contour map was prepared from the colour
fluorescence images.
A mouse fibroblast stained for DNA
(blue), transfected with SC35-GFP (a fluorescently
tagged RNA splicing factor) and microinjected with 40 nm
beads. A contour map was prepared from the colour
fluorescence images.